# Slightly overlapping data
set.seed(55)
df_cmp <- data.frame(
x1 = c(rnorm(50, 2, 1), rnorm(50, 4.5, 1)),
x2 = c(rnorm(50, 2, 1), rnorm(50, 4.5, 1)),
group = factor(rep(c("A", "B"), each = 50))
)
grid_cmp <- expand.grid(
x1 = seq(-1, 8, length.out = 150),
x2 = seq(-1, 8, length.out = 150)
)
par(mfrow = c(1, 2), mar = c(4, 4, 3, 1))
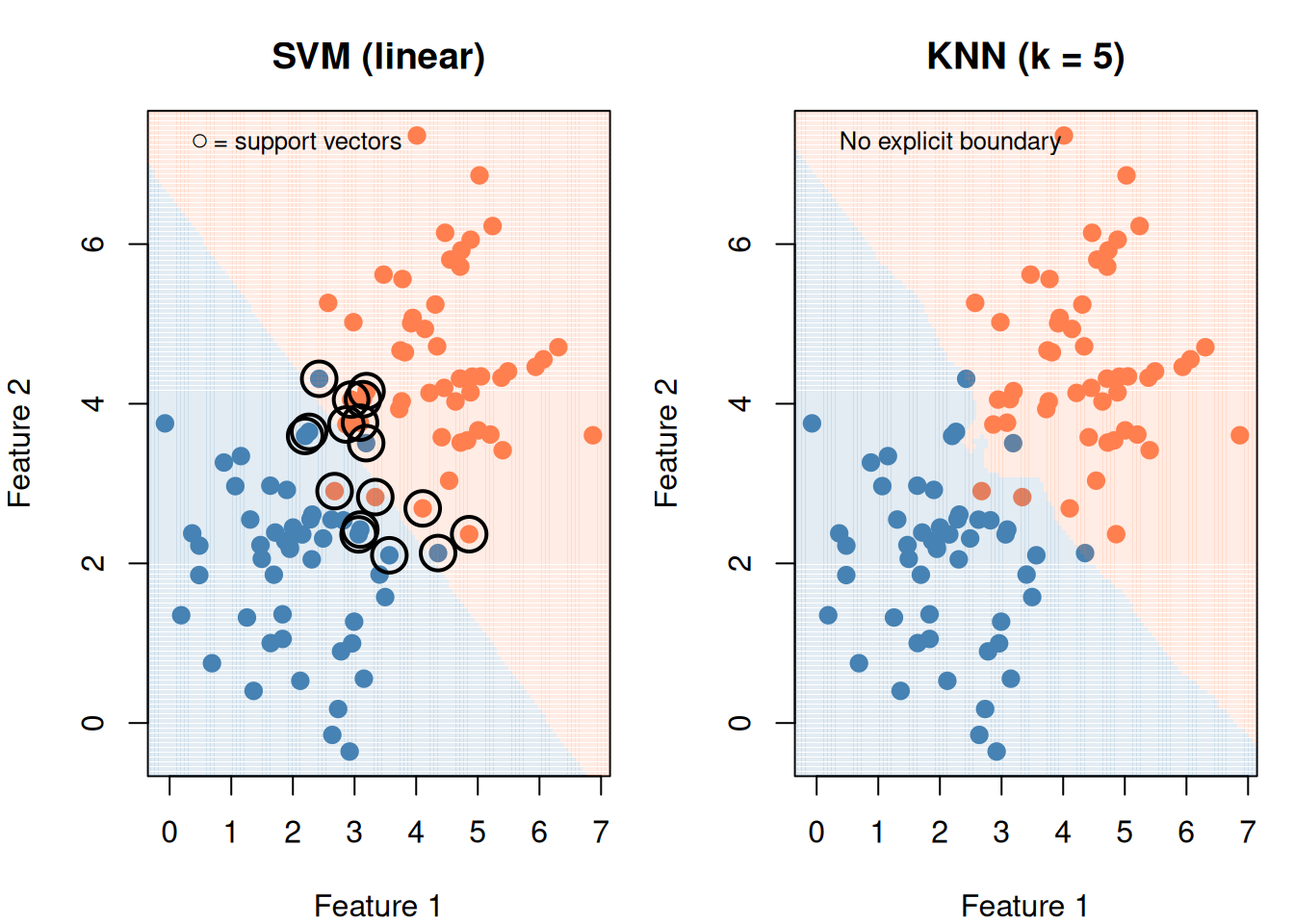
# SVM
fit_svm <- svm(group ~ x1 + x2, data = df_cmp, kernel = "linear", cost = 1)
grid_cmp$svm_pred <- predict(fit_svm, grid_cmp)
plot(df_cmp$x1, df_cmp$x2,
col = ifelse(df_cmp$group == "A", "steelblue", "coral"),
pch = 19, cex = 1.2, main = "SVM (linear)",
xlab = "Feature 1", ylab = "Feature 2")
points(grid_cmp$x1, grid_cmp$x2,
col = ifelse(grid_cmp$svm_pred == "A",
adjustcolor("steelblue", 0.15), adjustcolor("coral", 0.15)),
pch = 15, cex = 0.3)
points(df_cmp$x1[fit_svm$index], df_cmp$x2[fit_svm$index],
pch = 1, cex = 2.5, lwd = 2)
legend("topleft", bty = "n", legend = "○ = support vectors", cex = 0.8)
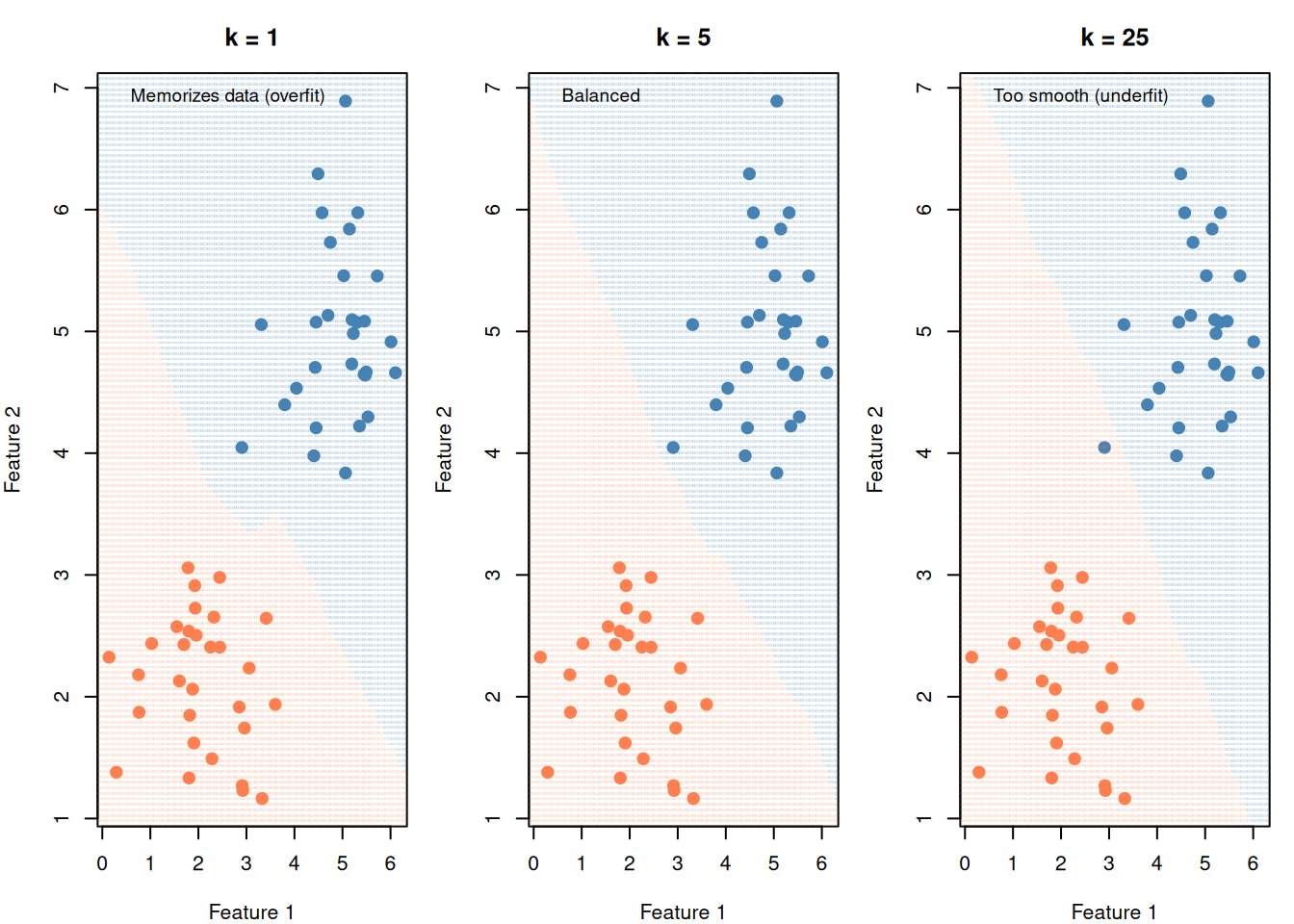
# KNN
knn_pred <- knn(
train = df_cmp[, c("x1", "x2")],
test = grid_cmp[, c("x1", "x2")],
cl = df_cmp$group, k = 5
)
plot(df_cmp$x1, df_cmp$x2,
col = ifelse(df_cmp$group == "A", "steelblue", "coral"),
pch = 19, cex = 1.2, main = "KNN (k = 5)",
xlab = "Feature 1", ylab = "Feature 2")
points(grid_cmp$x1, grid_cmp$x2,
col = ifelse(knn_pred == "A",
adjustcolor("steelblue", 0.15), adjustcolor("coral", 0.15)),
pch = 15, cex = 0.3)
legend("topleft", bty = "n", legend = "No explicit boundary", cex = 0.8)
par(mfrow = c(1, 1))